Background
During my time as a Chemistry undergraduate, I found myself most interested in organic macromolecular structure and reactions (specifically protein structure and enzyme catalysis). My dissertation project was to create an online resource (teaching NMR to undergraduate students) way back in the pre-Google days. The idea of combining protein structure with programming and web design led me to a Bioinformatics PhD in protein structure/classification in Prof. Christine Orengo's group at UCL.
After my PhD, I stayed on in the Orengo group as a post-doctoral researcher to help maintain and develop the group's main resource: the CATH database. I then decided to have a break from academia and took up a position as the Technical Director of a web development company. After a couple of years of commercial web/database/software development I was invited to consider a research post back in UCL. I realised I actually quite missed research so came back to UCL.
- 2016-present
- Principal Research Associate / Group Manager Orengo Group / Dept of Structural Molecular Biology / University College London (UCL)
- 2006-2016
- Research Fellow / Technical Manager Orengo Group / Dept of Structural Molecular Biology / University College London (UCL)
- 2002-2006
- Technical Director Arcray Webmedia
- 1998-2002
- PhD "Consensus Templates for Protein Structure Recognition" Dept of Biochemistry / University College London (UCL)
- 1994-1998
- MChem Sheffield University
Current Projects
CATH Protein Structure Classification Database
Genome3D Annotating Genomes with Structures

Research Interests
Consensus Structural Templates
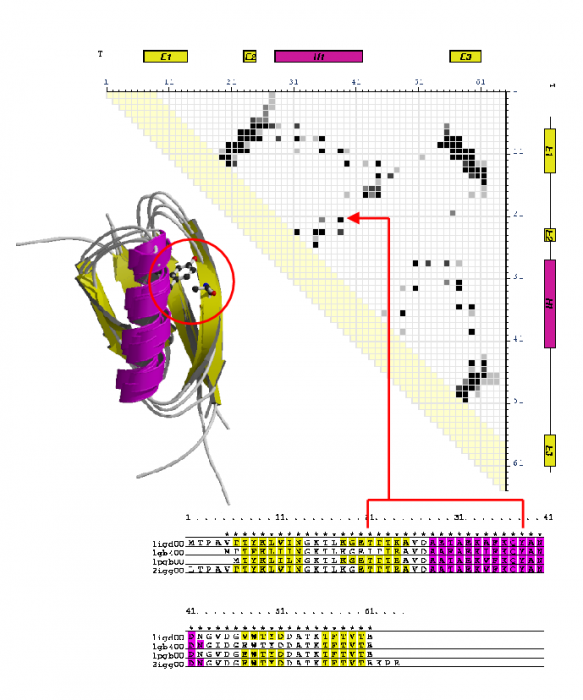
During my PhD, I looked at the classification, analysis and recognition of protein structures using the CATH protein structure classification database. I measured structural similarity by comparing contact maps: defined as points of contact (<8Å) between amino acid residues within a protein. By examining structures that were known to be related, it was possible to identify inter-residue contacts that were highly conserved during the process of evolution.
Part of this project involved deriving a protocol for generating accurate multiple structural alignments and 3D templates based on consensus contact patterns found in these alignments (see figure above). Templates were generated for all homologous superfamilies in CATH to create a library of unique and identifying 'fingerprint' patterns.
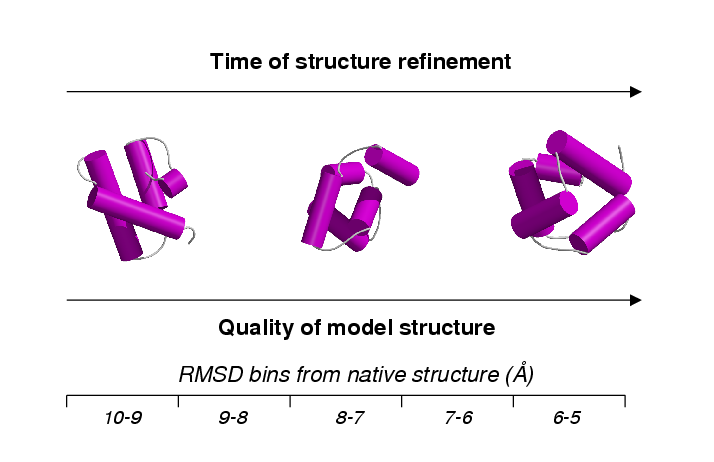
These templates were applied to a number of areas including the recognition of models generated at an early stage of ab initio protein structure prediction. Scanning these early models against a library of templates describing conserved contacts allowed the most likely superfamilies to be indentified. I then wrote an algorithm that performed fold recognition using only a limited set of contacts with the purpose of application to the early stages of experimental NMR structure determination. The multiple structural alignments were also used to generate a library of hidden markov models (HMMs). These structure-based sequence profiles were thoroughly benchmarked using a strict dataset of remote homologues and added sensitivity to existing sequence scans.
Other Interests
Jitsu / Judo
During my PhD, I started training regularly in TJF Jitsu (a self-defense martial art based on Shorinji Kan Jiu Jitsu). I became a instructor in 2005 and have taught at several clubs over the years - mainly ULU and Westminster. I graded to sandan (third degree black belt) in 2016 and still train and teach when I can.
I also train at Tsukuru Judo Academy - I'm hoping to find time to train and complete my black belt in 2014 2015 2016 2017 2018.
Music
I've played piano since I was young: trained classically for years, then been in a few different bands through school, Uni and here in London - a fairly eclectic mix of funk / soul / blues / jazz (ish). I do still get wheeled out for the occasional wedding or one-off gig. The most recent one involved going up and down the Thames on the paddle steamer "Dixie Queen" which was a lot of fun.

